Features
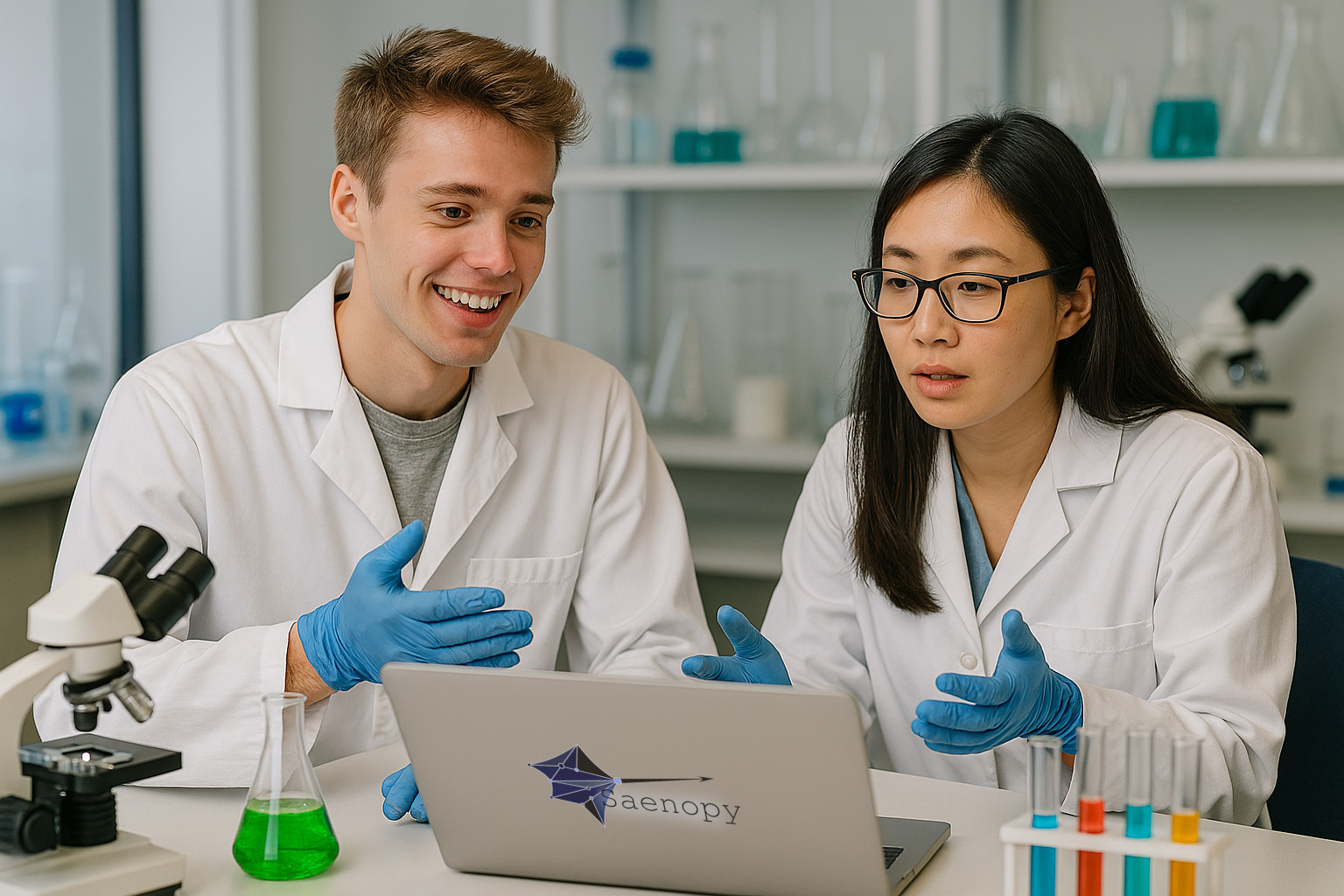
Saenopy provides powerful tools for cell mechanics research.
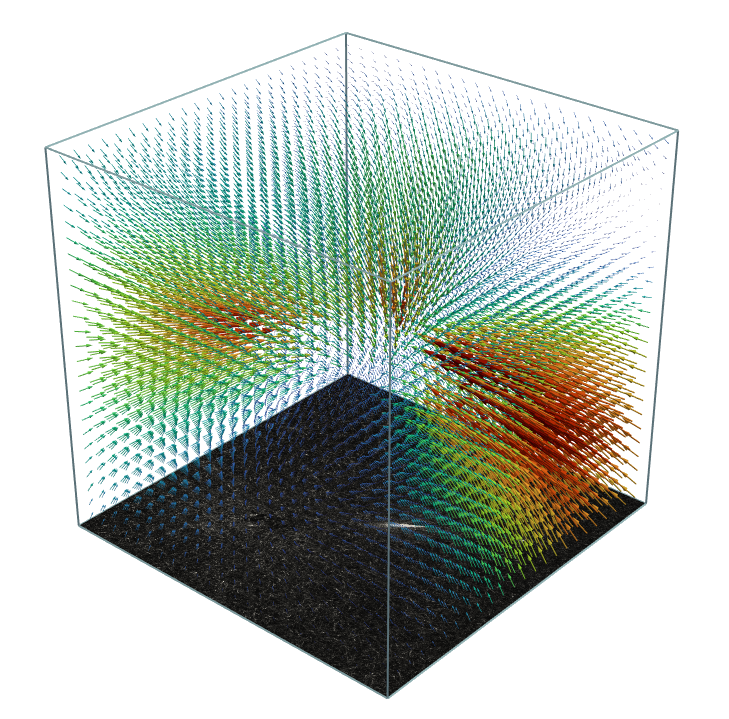
Calculate traction forces in three dimensions with high accuracy.
Support for various non-linear material models for accurate cell mechanics analysis.
Comprehensive visualization tools to analyze and present your results.
Fully scriptable with a comprehensive Python API for automation and integration.
User-friendly graphical interface for researchers without programming experience.
100% open source and free to use, modify, and distribute under MIT license.
Integrations
Aside from Saenopy's main use for 3D traction force microscopy, we provide integrations to related methods to assess cellular forces.
Analyze forces in multicellular aggregates (so-called spheroids) embedded in 3D biopolymer networks.
Mark C., Grundy T., Strissel P., et al. (2020). "Collective forces of tumor spheroids in three-dimensional biopolymer networks". In eLife 9:e51912.
Use fiber alignment as a proxy for force when material properties are not available.
Böhringer D., Bauer A., Moravec I., et al. (2023). "Fiber alignment in 3D collagen networks as a biophysical marker for cell contractility". Matrix Biology, 124, pp.39-48.
Meet the Team
The brilliant minds behind Saenopy and its related projects.
Creator of Saenopy, pylustrator, and cameratransform. Specializes in scientific software development and data visualization.
Physicist focused on time series analysis and complex systems. Creator of jointforces for analyzing multicellular aggregates.
Support Development
Saenopy is 100% open source. Your donations help us continue development and add new features.
Your support helps us continue to develop and maintain Saenopy.